In silico screening of azo acetohydrazide derivatives as potential antidiabetic agents through alpha glucosidase inhibition
DOI:
https://doi.org/10.29303/aca.v8i2.263Keywords:
Type 2 Diabetes Mellitus, Alpha Glucosidase, Azo-Acetohydrazide,, in Silico StudyAbstract
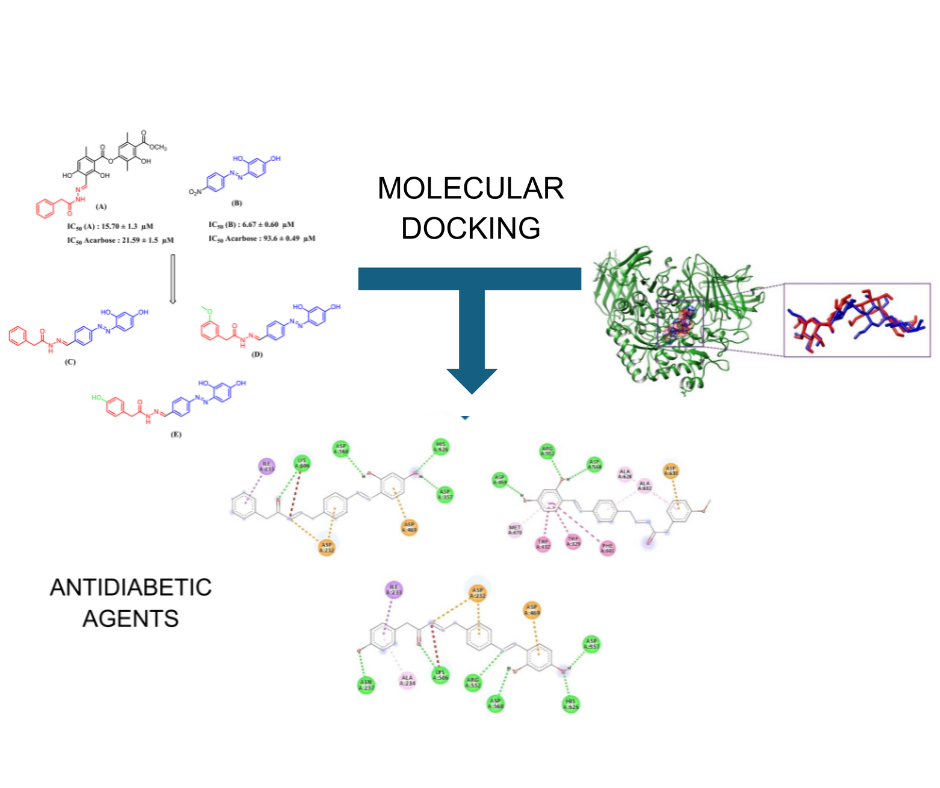
Type 2 diabetes mellitus is a progressive metabolic disorder marked by persistent hyperglycemia resulting from insulin resistance, insufficient insulin secretion, or a combination of both conditions. A potential therapeutic target for treating this illness is the suppression of the alpha-glucosidase enzyme, which is a key regulator of postprandial glucose absorption and carbohydrate metabolism. This work conducted a thorough in silico analysis to evaluate the efficacy of three synthesised azo-acetohydrazide derivatives (designated compounds C, D, and E) as alpha-glucosidase inhibitors. Pharmacokinetic properties and drug-likeness characteristics were evaluated using the SwissADME platform, while molecular docking simulations were conducted with AutoDockTools against the alpha-glucosidase enzyme structure sourced from the Protein Data Bank (PDB ID: 3W37), utilising acarbose as a reference inhibitor. All three compounds adhered to essential drug-likeness criteria, including those established by Lipinski, Veber, and Ghose, and exhibited advantageous physicochemical characteristics, such as appropriate molecular weight, lipophilicity, topological polar surface area, and aqueous solubility. The compounds were anticipated to have significant gastrointestinal absorption and had no ability to penetrate the blood-brain barrier, suggesting a minimal risk of central nervous system damage. Compound E demonstrated the highest binding affinity among the compounds, with a docking score of –7.80 kcal/mol, and formed multiple stabilising interactions within the enzyme's active region, such as hydrogen bonds, hydrophobic contacts, and electrostatic interactions. This study's novelty is the computational discovery of azo-acetohydrazide scaffolds as inadequately investigated chemical entities with potential antidiabetic properties. These findings establish a theoretical basis for continued development and substantiate future endeavours in the synthesis and experimental validation of these molecules. This study validates the efficacy of structure-based drug design in discovering novel alpha glucosidase inhibitors and identifies compound E as a good candidate for further in vitro and in vivo research in type 2 diabetes treatment.
Downloads
References
Sakamoto-Hojo, E. T. (2022). Mechanisms underlying the pathophysiology of type 2 diabetes: From risk factors to oxidative stress, metabolic dysfunction, and hyperglycemia. Mutation Research/Genetic Toxicology and Environmental Mutagenesis, 874–875, 503437.
IDF Diabetes Atlas 2025 | Global Diabetes Data & Insights. (2025). Retrieved July 11, 2025, from https://diabetesatlas.org/resources/idf-diabetes-atlas-2025/
Lu, H., Xie, T., Wu, Q., Hu, Z., Luo, Y., & Luo, F. (2023). Alpha-Glucosidase Inhibitory Peptides: Sources, Preparations, Identifications, and Action Mechanisms. Nutrients, 15(19), 1–24.
Kashtoh, H., & Baek, K. H. (2022). Recent Updates on Phytoconstituent Alpha-Glucosidase Inhibitors: An Approach towards the Treatment of Type Two Diabetes. Plants, 11(20), 2722.
Chen, Y., Xiao, Y., Lian, G., Yi, J., & Liu, X. (2023). Pneumatosis intestinalis associated with α-glucosidase inhibitors: a pharmacovigilance study of the FDA adverse event reporting system from 2004 to 2022. 1–10. Expert Opinion on Drug Safety.
Sharma, P., Sharma, V., Mathpal, S., Tewari, D., Chandra, S., & Tamta, S. (2024). Identification of alpha-glucosidase enzyme inhibitors from phytochemicals via integrated deep learning, molecular docking, molecular dynamics simulation, and MMPBSA analysis. South African Journal of Botany, 167, 48–61.
Banaszak-Leonard, E., Fayeulle, A., Franche, A., Sagadevan, S., & Billamboz, M. (2021). Antimicrobial azo molecules: a review. Journal of the Iranian Chemical Society, 18(11), 2829–2851.
Di Martino, M., Sessa, L., Di Matteo, M., Panunzi, B., Piotto, S., & Concilio, S. (2022). Azobenzene as Antimicrobial Molecules. Molecules, 27(17), 5643.
Dandge, S. V., Nikume, S. R., & Bendre, R. S. (2024). An efficient synthesis, characterisation, antimicrobial and anticancer activities of azo dyes derived from eugenol. Synthetic Communications, 54(4), 282–292.
Tadić, J. D., Lađarević, J. M., Vitnik, Ž. J., Vitnik, V. D., Stanojković, T. P., Matić, I. Z., & Mijin, D. (2021). Novel azo pyridone dyes based on dihydropyrimidinone skeleton: Synthesis, DFT study and anticancer activity. Dyes and Pigments, 187, 109123.
Mezgebe, K., & Mulugeta, E. (2022). Synthesis and pharmacological activities of azo dye derivatives incorporating heterocyclic scaffolds: a review. RSC Advances, 12(40), 25932–25946.
[Agili, F. (2024). Novel Hydrazide Hydrazone Derivatives as Antimicrobial Agents: Design, Synthesis, and Molecular Dynamics. Processes, 12(6), 1055.
Ur Rahman, S., Alam, A., Parveen, Z., Zainab, Assad, M., Adnan Ali Shah, S., Rafiq, H., Ayaz, M., Latif, A., Naveed Umar, M., Ali, M., & Ahmad, M. (2024). Novel acyl hydrazide derivatives of polyhydroquinoline as potent antidiabetic and anti-glycating agents: Synthesis, in vitro α-amylase, α-glucosidase inhibition and anti-glycating activity with molecular docking insights. Bioorganic Chemistry, 150, 107501.
Valipour, M., Zakeri, Z., Kiadaliry, K., Mojtabavi, S., Faramarzi, M. A., Sayyad, M. S., Seyedabadi, M., Ghasemian, M., Hashemi, S. M., & Irannejad, H. (2024). Design, synthesis, α-glucosidase inhibition and hypoglycemic activity of 3-aceto(benzo)hydrazide-1,2,4-triazines as potential antidiabetic agents. European Journal of Medicinal Chemistry Reports, 12, 100207.
Han, M. İ., Yeşil Baysal, Ö. D., Başaran, G. Ş., Sezer, G., Telci, D., & Küçükgüzel, G. G. (2022). Design, synthesis and anticancer activity studies of novel 4-butylaminophenyl hydrazide-hydrazones as apoptotic inducers. Tetrahedron, 115, 132797.
Rana, M., Cho, H. J., Arya, H., Bhatt, T. K., Bhar, K., Bhatt, S., … Sharma, A. K. (2022). Azo-Stilbene and Pyridine-Amine Hybrid Multifunctional Molecules to Target Metal-Mediated Neurotoxicity and Amyloid-β Aggregation in Alzheimer’s Disease. Inorganic Chemistry, 61(27), 10294–10309.
Li, Y., Sui, J., Cui, L. S., & Jiang, H. L. (2023). Hydrogen Bonding Regulated Flexibility and Disorder in Hydrazone-Linked Covalent Organic Frameworks. Journal of the American Chemical Society, 145(2), 1359–1366.
[Daina, A., Michielin, O., & Zoete, V. (2017). SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Scientific Reports, 7(March), 1–13.
Angelova, V. T., Pencheva, T., Vassilev, N., K-Yovkova, E., Mihaylova, R., Petrov, B., & Valcheva, V. (2022). Development of New Antimycobacterial Sulfonyl Hydrazones and 4-Methyl-1,2,3-thiadiazole-Based Hydrazone Derivatives. Antibiotics, 11(5), 1–22.
Silva, A. M., Martins-Gomes, C., Ferreira, S. S., Souto, E. B., & Andreani, T. (2022). Molecular Physicochemical Properties of Selected Pesticides as Predictive Factors for Oxidative Stress and Apoptosis-Dependent Cell Death in Caco-2 and HepG2 Cells. International Journal of Molecular Sciences, 23(15).
Wu, D., Chen, Q., Chen, X., Han, F., Chen, Z., & Wang, Y. (2023). The blood–brain barrier: structure, regulation, and drug delivery. Signal Transduction and Targeted Therapy, 8(1).
Zhao, M., Ma, J., Li, M., Zhang, Y., Jiang, B., Zhao, X., … Qin, S. (2021). Cytochrome p450 enzymes and drug metabolism in humans. International Journal of Molecular Sciences, 22(23), 1–16.
Ramírez, D., & Caballero, J. (2018). Is It Reliable to Take the Molecular Docking Top Scoring Position as the Best Solution without Considering Available Structural Data?. Molecules, 23(5).
Tagami, T., Yamashita, K., Okuyama, M., Mori, H., Yao, M., & Kimura, A. (2013). Molecular basis for the recognition of long-chain substrates by plant α-glucosidases. Journal of Biological Chemistry, 288(26), 19296–19303.
Menacer, R., Bouchekioua, S., Meliani, S., & Belattar, N. (2024). New combined Inverse-QSAR and molecular docking method for scaffold-based drug discovery. Computers in Biology and Medicine, 180, 108992.
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2025 Muhammad Zulqurnain, Sakti Hidayati Fikriya, Andi Suharman, Annisha Noor Dienna, First Ambar Wati

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Authors who publish with ACA: Acta Chimica Asiana agree to the following terms:
- Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. This license allows authors to use all articles, data sets, graphics, and appendices in data mining applications, search engines, web sites, blogs, and other platforms by providing an appropriate reference. The journal allows the author(s) to hold the copyright without restrictions and will retain publishing rights without restrictions.
- Authors are able to enter into separate, additional contractual arrangements for the non-exclusive distribution of the journal's published version of the work (e.g., post it to an institutional repository or publish it in a book), with an acknowledgement of its initial publication in ACA: Acta Chimica Asiana.
- Authors are permitted and encouraged to post their work online (e.g., in institutional repositories or on their website) prior to and during the submission process, as it can lead to productive exchanges, as well as earlier and greater citation of published work (See The Effect of Open Access).





 Indonesian Chemical Society, Chapter Nusa Tenggara. Jalan Majapahit 62 Mataram, University of Mataram, 83125, Indonesia
Indonesian Chemical Society, Chapter Nusa Tenggara. Jalan Majapahit 62 Mataram, University of Mataram, 83125, Indonesia
